Microecology Close to Home
Aaron Best, Ph.D. | Harrison C. and Mary L. Visscher Professor of Genetics
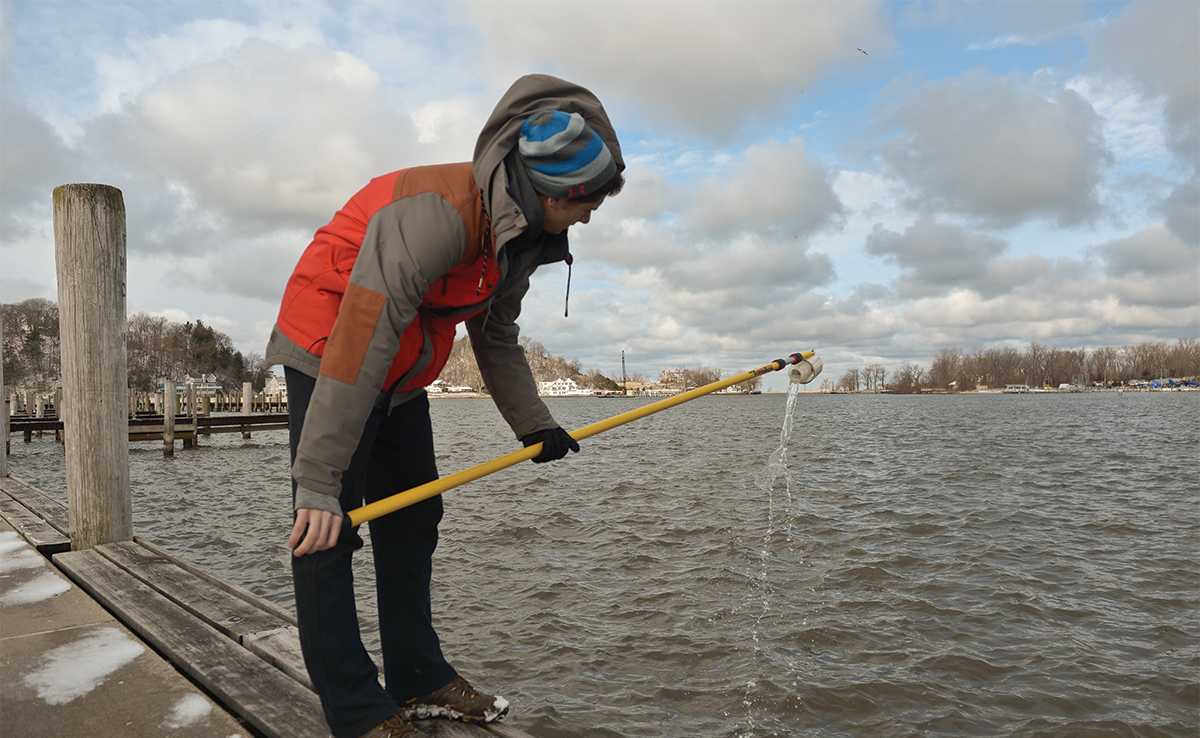
For more than 15 years, Hope faculty members have worked alongside students to research Lake Macatawa’s watershed. Decades of agricultural run-off polluted it with nutrients that can kill fish, harm plant life, lead to algal blooms, and trigger high levels of E. coli bacteria.
To spark students’ interest in saving local habitats, and to monitor nutrient levels and bacterial populations to determine whether current environmental restoration programs are effective in the Macatawa watershed, Hope College established the “Day1: Watershed” program in 2015. Starting with their first day on campus, science-focused freshmen can engage in hands-on research with biologist Dr. Aaron Best, chemist Dr. Brent Krueger and physicist Dr. Catherine Mader through a First Year Seminar and introductory biology lab.
Throughout the last three years, students have collected and analyzed water samples and contributed their findings to a large-scale research project that has four long-term goals: to offer the Holland community information about students’ latest investigations; to reduce the amount of nutrients in the lake; to decrease bacterial pollution; and, ultimately, to clarify Lake Macatawa. In December 2017 the Macatawa Area Coordinating Council honored the Day1: Watershed project with its Watershed Stakeholder of the Year Award, which honors efforts to improve the water quality of the watershed.

“We are gathering samples that monitor the amounts and types of bacteria present, including E. coli. As we begin to understand the levels of E. coli present, we are also trying to determine which other microbes exist in the watershed and how the levels of E. coli can be correlated with the levels of the other microbes,” Best explains.
“It will take time before significant change is noticed. We are currently monitoring for nutrients like phosphorus. The goal is to reduce those levels to 50 parts per billion of phosphorus, a 70 percent reduction over current levels.”
This research is the first of its kind, as students are chronicling multiple strains of E. coli that occur in unique environments — strains potentially unassociated with host organisms. The project is funded by Best’s three-year, $775,316 grant from the National Science Foundation, the largest single research grant ever received by a Hope professor.
In addition to Hope students, Best’s research team includes Dr. Matthew DeJongh, professor of computer science; Dr. Michael Pikaart, associate professor of chemistry; and Dr. Stephen Scogin, assistant professor of biology and education. Each member of the team brings particular expertise to the project in areas such as bioinformatics data analysis, water chemistry, and research into how professors can provide students the best learning experience.
To improve their watershed monitoring techniques, Best and his colleagues are considering alternatives to E. coli culture counts. They’re investigating whether other bacterial organisms, or sets of organisms, may indicate whether water is safe for human contact — and whether there are better ways to monitor them than the current process of counting. They also want to determine which types of genes are present in the E. coli genome itself, which strains of the bacteria are problematic, and how they can be identified in the first place.
“Almost all of the genome sequences of E. coli available in public databases are derived from a diseased state or a clinical source, and there are thousands of them,” Best explains. “Very few are derived from a non-diseased state. So, with that in mind, we’re trying to answer two questions: Are the E. coli present in Lake Macatawa likely to be pathogens? And, if so, what is the source?”
In summer 2017, Best worked with 21 Hope student research assistants. For roughly three months, they sampled 12 different sites at the watershed every Tuesday. One group of students studied the levels of chemicals, nutrients and sediments in the water. Another group tested water samples to determine how much E. coli was in the watershed; these samples were used to isolate individual strains, which were then used for genome sequencing and compared to other genomes. A third group studied the same water samples as the genome group, but isolated DNA from all of the organisms that were present; DNA was then sequenced for particular genes to identify each of the organisms.
“In doing so, you come up with community profiles. Which types of bacteria are present, and how much?” Best explains. “You then analyze those for patterns. Does the winter look the same as the summer? What about rain patterns?”
“There are clearly seasonal variations as well. We’re beginning to understand what is present, and what that might mean. There is a nice seasonal cycle at this point, as we can show what freshwater systems look like during the summer and winter months. We can then compare different years this way and see if patterns are consistent throughout.”
To date, the research has proven that sediment and nutrient levels are still very high, although they should decline over time. For example, phosphorus and nitrate levels are above target, and E. coli remains high in streams. Nonetheless, although the lake’s strains of E. coli are similar in appearance to traditional E. coli, they don’t have the same traits as known pathogens. The number of genes that help E. coli to cause disease appears to be much lower than in traditional strains.
Much of this research is integrated into Hope classes. Successive groups of students can learn about — and contribute to — the findings every year.
“Students from the Day1: Watershed program, as well as upper-level microbiology courses, have continued to work on the broader watershed project in my research lab since they already know about the science and techniques of the research from past experiences,” Best adds. “This greatly advances the speed of the project and allows us to analyze incoming data at a deeper level.”
Best expects that this research may ultimately add 750 to 800 new genome sequences to public databases, each of which can be used by researchers worldwide.
“As we continue to write about the initial data, we will not only be contributing to public databases, but we will also educate people locally and nationally about our findings and their impact,” he says. “We have already presented our work at community informational meetings as well as national conferences, and will continue to do so, as we learn more about the watershed — and strive to improve it.”